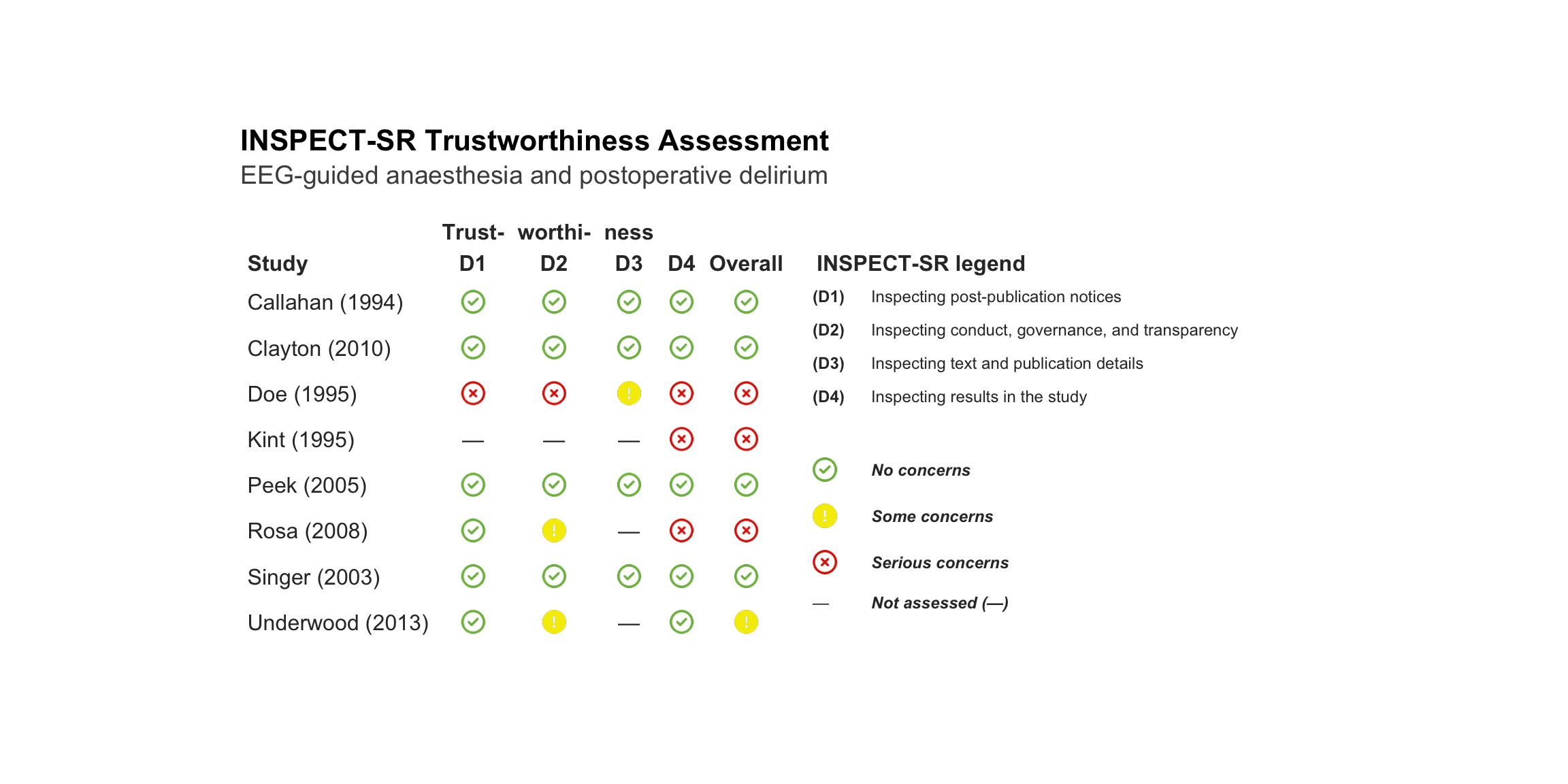
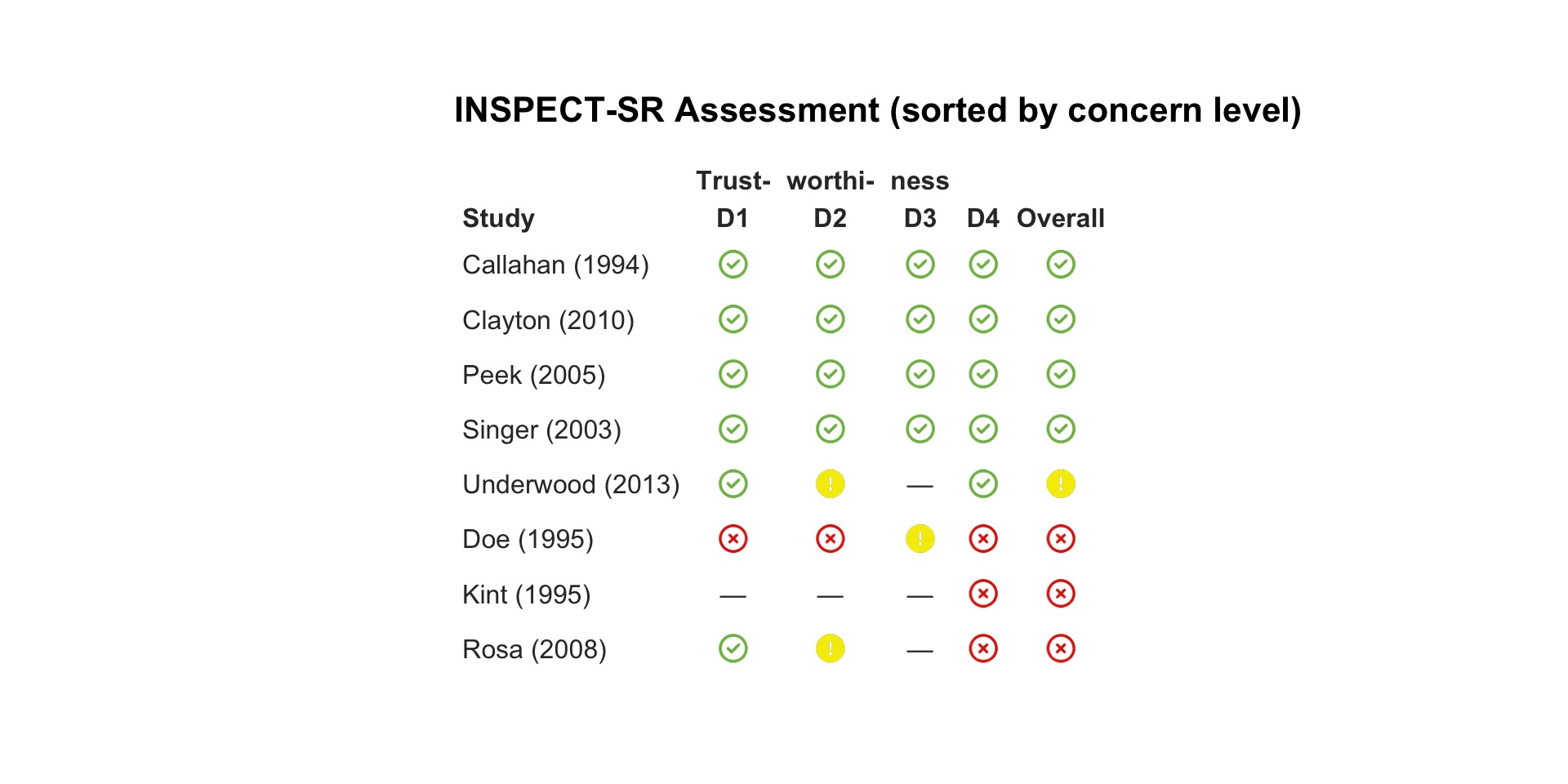
| Callahan (1994) |
Carlisle |
Baseline p-value distribution |
k = 3, fisher combined p = 0.3788, plausible |
Pass |
| GRIM |
ASA_score (Intervention) |
mean = 2.5, n = 46 |
Pass |
|
ASA_score (Control) |
mean = 2.5, n = 46 |
Pass |
| N-consistency |
Total randomised = Intervention + Control |
expected = 92, observed = 92 |
Pass |
|
Intervention: Randomised = Analysed + Lost |
expected = 46, observed = 46 |
Pass |
|
Control: Randomised = Analysed + Lost |
expected = 46, observed = 46 |
Pass |
|
Intervention: Lost <= Randomised |
expected = 46, observed = 3 |
Pass |
|
Control: Lost <= Randomised |
expected = 46, observed = 2 |
Pass |
| P-value |
Delirium incidence |
reported p = 0.27, recalculated p = 0.2674 (diff 0.002593) |
Pass |
| Clayton (2010) |
Carlisle |
Baseline p-value distribution |
k = 3, fisher combined p = 0.46, plausible |
Pass |
| N-consistency |
Total randomised = Intervention + Control |
expected = 202, observed = 202 |
Pass |
|
Intervention: Randomised = Analysed + Lost |
expected = 101, observed = 101 |
Pass |
|
Control: Randomised = Analysed + Lost |
expected = 101, observed = 101 |
Pass |
|
Intervention: Lost <= Randomised |
expected = 101, observed = 3 |
Pass |
|
Control: Lost <= Randomised |
expected = 101, observed = 5 |
Pass |
| P-value |
Delirium incidence |
reported p = 0.1, recalculated p = 0.09545 (diff 0.004552) |
Pass |
| Doe (1995) |
Carlisle |
Baseline p-value distribution |
k = 4, fisher combined p = 0.05859, plausible |
Pass |
| GRIM |
ASA_score (Intervention) |
mean = 2.45, n = 90 |
Fail |
|
ASA_score (Control) |
mean = 2.55, n = 90 |
Fail |
|
Pain_VAS (Intervention) |
mean = 4.75, n = 90 |
Fail |
|
Pain_VAS (Control) |
mean = 4.65, n = 90 |
Fail |
| N-consistency |
Total randomised = Intervention + Control |
expected = 180, observed = 180 |
Pass |
|
Intervention: Randomised = Analysed + Lost |
expected = 90, observed = 90 |
Pass |
|
Control: Randomised = Analysed + Lost |
expected = 90, observed = 90 |
Pass |
|
Intervention: Lost <= Randomised |
expected = 90, observed = 2 |
Pass |
|
Control: Lost <= Randomised |
expected = 90, observed = 3 |
Pass |
| P-value |
Delirium incidence |
reported p = 0.001, recalculated p = 0.000407 (diff 0.000593) |
Pass |
|
Delirium duration |
reported p = 0.002, recalculated p = 0.001582 (diff 0.0004176) |
Pass |
| Kint (1995) |
Carlisle |
Baseline p-value distribution |
k = 3, fisher combined p = 0.4555, plausible |
Pass |
| GRIM |
Duration_surgery_min (Intervention) |
mean = 175, n = 150 |
Pass |
|
Duration_surgery_min (Control) |
mean = 180, n = 150 |
Pass |
| N-consistency |
Total randomised = Intervention + Control |
expected = 300, observed = 300 |
Pass |
|
Intervention: Randomised = Analysed + Lost |
expected = 150, observed = 147 |
Fail |
|
Control: Randomised = Analysed + Lost |
expected = 150, observed = 146 |
Fail |
|
Intervention: Lost <= Randomised |
expected = 150, observed = 5 |
Pass |
|
Control: Lost <= Randomised |
expected = 150, observed = 8 |
Pass |
| P-value |
Delirium incidence |
reported p = 0.08, recalculated p = 0.07734 (diff 0.002663) |
Pass |
| Peek (2005) |
Carlisle |
Baseline p-value distribution |
k = 4, fisher combined p = 0.3872, plausible |
Pass |
| GRIM |
Duration_surgery_min (Intervention) |
mean = 185, n = 230 |
Pass |
|
Duration_surgery_min (Control) |
mean = 190, n = 230 |
Pass |
| N-consistency |
Total randomised = Intervention + Control |
expected = 460, observed = 460 |
Pass |
|
Intervention: Randomised = Analysed + Lost |
expected = 230, observed = 230 |
Pass |
|
Control: Randomised = Analysed + Lost |
expected = 230, observed = 230 |
Pass |
|
Intervention: Lost <= Randomised |
expected = 230, observed = 9 |
Pass |
|
Control: Lost <= Randomised |
expected = 230, observed = 12 |
Pass |
| P-value |
Delirium incidence |
reported p = 0.04, recalculated p = 0.04238 (diff 0.002379) |
Pass |
|
Duration of delirium |
reported p = 0.4, recalculated p = 0.3958 (diff 0.004209) |
Pass |
| Rosa (2008) |
Carlisle |
Baseline p-value distribution |
k = 6, fisher combined p = 4.265e-05, too_similar |
Fail |
| N-consistency |
Total randomised = Intervention + Control |
expected = 240, observed = 240 |
Pass |
|
Intervention: Randomised = Analysed + Lost |
expected = 120, observed = 120 |
Pass |
|
Control: Randomised = Analysed + Lost |
expected = 120, observed = 120 |
Pass |
|
Intervention: Lost <= Randomised |
expected = 120, observed = 2 |
Pass |
|
Control: Lost <= Randomised |
expected = 120, observed = 3 |
Pass |
| P-value |
Delirium incidence |
reported p = 0.005, recalculated p = 0.00511 (diff 0.0001103) |
Pass |
| Singer (2003) |
Carlisle |
Baseline p-value distribution |
k = 3, fisher combined p = 0.4289, plausible |
Pass |
| N-consistency |
Total randomised = Intervention + Control |
expected = 1232, observed = 1232 |
Pass |
|
Intervention: Randomised = Analysed + Lost |
expected = 614, observed = 614 |
Pass |
|
Control: Randomised = Analysed + Lost |
expected = 618, observed = 618 |
Pass |
|
Intervention: Lost <= Randomised |
expected = 614, observed = 16 |
Pass |
|
Control: Lost <= Randomised |
expected = 618, observed = 13 |
Pass |
| P-value |
Delirium incidence |
reported p = 0.67, recalculated p = 0.6714 (diff 0.001373) |
Pass |
| Underwood (2013) |
Carlisle |
Baseline p-value distribution |
k = 3, fisher combined p = 0.4773, plausible |
Pass |
| GRIM |
Duration_anaesthesia_min (Intervention) |
mean = 210, n = 80 |
Pass |
|
Duration_anaesthesia_min (Control) |
mean = 198, n = 80 |
Pass |
| N-consistency |
Total randomised = Intervention + Control |
expected = 160, observed = 160 |
Pass |
|
Intervention: Randomised = Analysed + Lost |
expected = 80, observed = 80 |
Pass |
|
Control: Randomised = Analysed + Lost |
expected = 80, observed = 80 |
Pass |
|
Intervention: Lost <= Randomised |
expected = 80, observed = 4 |
Pass |
|
Control: Lost <= Randomised |
expected = 80, observed = 6 |
Pass |
| P-value |
Delirium incidence |
reported p = 0.02, recalculated p = 0.02506 (diff 0.005056) |
Pass |